 Charles Marchini joined the program on January 1, 2021, while a first-year bioengineering student, and is expected to graduate in 2025. He completed his undergraduate degree in electrical engineering at Northern Illinois University (NIU), and in 2020 NIU awarded him with Upper Division Honors. He is completing lab rotations under the advisement of Brad Sutton and Martha Gillette. He has completed coursework in neuroscience and scientific professionalism as well as training in writing for publication, in addition to bioengineering coursework. In May 2022, he presented his research at the annual conference for the International Society for Magnetic Resonance in Medicine.
Charles Marchini joined the program on January 1, 2021, while a first-year bioengineering student, and is expected to graduate in 2025. He completed his undergraduate degree in electrical engineering at Northern Illinois University (NIU), and in 2020 NIU awarded him with Upper Division Honors. He is completing lab rotations under the advisement of Brad Sutton and Martha Gillette. He has completed coursework in neuroscience and scientific professionalism as well as training in writing for publication, in addition to bioengineering coursework. In May 2022, he presented his research at the annual conference for the International Society for Magnetic Resonance in Medicine.
Marchini has participated in the Special Topics in MBM course as well as other trainee activities such as the annual MBM Retreat, the Summer Journal Club readings/discussion meetings, Frontiers in Miniature Brain Machinery lectures, and multiple outreach efforts.
Marchini’s tutorial on “Data Collection in MRI Scanners,” December 2021
Chiefly among those outreach efforts are his development of video tutorials to help the public better understand topics related to imaging: “Data Collection in MRI Scanners,” “How to Shift Functions,” and “Arterial Spin Labeling MRI.”
Marchini’s tutorial on “How to Shift Functions,” April 2022
In addition, he taught a live lesson on covariance with fellow trainee Olawale Salaudeen to 62 middle-school students at Franklin STEAM Academy over 3 class periods on April 21, 2022.
“I learned a lot during the 3 lessons I did on covariance,” Charles explains. “I think the animation I made was helpful in getting me to think about science communication with high-school-level math. It was my first time teaching middle schoolers, and I was glad to see them asking questions, responding to questions, and understanding the concepts.”
He also participated in the MBM Program’s booth at the Urbana Farmer’s Market in June 2022, part of a larger effort with the University of Illinois to provide “Science at the Market” and share simple science lessons with the public.

In 2023, he plans to pursue his external research experience with the FDA’s MR safety division to learn about MR safety, implants, and to reinforce MR physics principles. He will primarily be working on the safety of implanted devices and MRI, such as stimulators and their electrodes, and learning how to simulate and measure effects.
Research Highlights (in his own words):
My research project for the MBM program is on implementing MRI perfusion quantification imaging using a partial separability method and arterial spin labeling (ASL). MRI perfusion quantification imaging using ASL is an MRI technique that does not require exogenous contrast agents and can image perfusion (blood interacting with tissue) in units of milliliters of blood per grams of brain tissue per minute. The major downfall with this technique is a very low signal-to-noise ratio compared to other MRI techniques. This is because the blood only makes up about 2% of a typical MRI image measuring water density, as is done with ASL.
Marchini’s tutorial on “Arterial Spin Labeling,” November 2022
ASL may also be used for functional MRI (fMRI). Changes in blood flow across time to a region of the brain can be measured using ASL. The current downfalls of this method are the low temporal and spatial resolution. Each ASL image can take 3 seconds to acquire, which limits the number of samples that can be acquired per repetition, meaning a decreased spatial resolution. The Partial Separability (PS) model can be implemented to increase the spatial and temporal resolution of ASL and ASLfMRI.
The PS method is a mathematical technique that is used to construct an MRI image using a small amount of data relative to the standard technique. The standard technique method requires that many data points be acquired during signal acquisition at the MRI scanner because the data are fully sampled. Each fully sampled set of data is used to reconstruct the image at that time point. If multiple time points are acquired, and there is a spatiotemporal correlation within the time points, the data can be broken down and compressed into a spatial basis and a temporal basis consisting of less data than the fully sampled data. Therefore, less sampling is required for each time point so each image is faster to acquire at the scanner. When the data is fully sampled, it takes longer to acquire all the data and the signal decays quickly, so the data taken has low signal. In the case of the standard reconstruction, the signal is decayed when we are collecting the high frequency components (such as edges, see Fourier analysis) of the signal. With the signal acquisition corresponding to the PS method, we can collect higher frequency data before the signal has decayed.

The most prominent parts of the spatial basis and the temporal basis are selected for (re)construction of the images. By selecting the most prominent parts, some of the noise (undesired signal) is filtered out. Other desired parts of the signal may also be filtered out. If the noise is reduced more than the desired signal, an increase in the signal-to-noise ratio and image quality results.
To apply the PS model to MRI perfusion, a reconstruction algorithm was used in Matlab. The input into the algorithm is the data file which includes the raw data from the MRI scanner. The temporal basis is found by doing a singular value decomposition of a high contrast part of the raw data. The singular value decomposition reorganizes the data so the functions representing how the brain volumes change through time can be extracted. The high-contrast part of the data is at the center of k-space (raw data in the spatial frequency domain), where there is low spatial resolution but high signal. The spatial basis is found by solving for the spatial basis that causes the reconstructed images to match the partially sampled data across all time points and conforms to the low rank constraint (so a limited number of spatial and temporal basis functions are used which is part of the partial separability model). This differs from the standard reconstruction method which performs a Fourier transform to the fully sampled imaging data for each time point separately. The hypothesis of this project is that the spatiotemporal resolution of ASL MRI can be improved by incorporating knowledge of the low rank property of the time series into the reconstruction.
Previously, we have shown that PSASL allows for a reconstruction with sharper images in the superior-inferior direction than the standard reconstruction. This semester, we have demonstrated that the PS model can also allow for better ASLfMRI.
Marchini’s presentation at the 2022 MBM Retreat on “MRI Perfusion Imaging with Partial Separability”
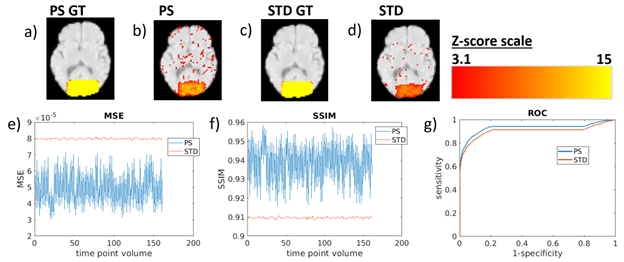
Increased blood flow was detected with a higher sensitivity and specificity for the PS reconstruction, demonstrated by a simulation (Figure 1). A human brain was simulated with an increase in blood flow for a 20 second on 20 second off task and input into the reconstruction algorithms. The mean squared error between the ground truth (the input into the simulation) and the reconstruction was lower for the PS recon. The structural similarity index was also higher for the PS recon (higher SSIM means more similarity between the ground truth and the reconstructed image). The specificity and sensitivity was also higher for the PS reconstruction, as shown by the area under the curve of the receiver operating characteristic graph, meaning more statistically significant results for showing increased blood flow.
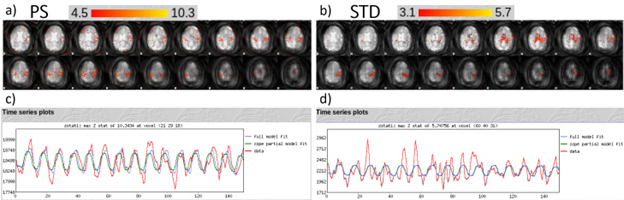
On a 3T scanner, an ASLfMRI finger tapping task was performed (Figure 2). The acquisition consisted of 32 coronal slices and 64 by 64 in plane pixels. The task consisted of 20 seconds off 20 seconds on of finger tapping for 648 seconds., and blood flow was detected as expected with real-world human data.
The results from this past semester have been submitted to an ISMRM conference and will likely be presented in May. After some more progress on this project is made, a paper will be drafted and submitted to a journal for publication. For future work on this project, we plan on obtaining ASLfMRI scans with a higher spatial resolution than would be possible with the standard acquisition at the same temporal resolution. We also have ideas for other projects such as applying this method at 7T and imaging flow through organoids.
I will be visiting the FDA in the Spring semester to work more with MRI electromagnetics and MRI implant safety, which will help me learn more about the regulations and potentially how to understand new safety procedures related to 7T MRI scans.

